Science
Tackling Cancer’s Loss of Function Mutations
Tremendous progress is being made in our understanding and treatment of cancer. Different biologic and therapeutic approaches have resulted in significantly improved clinical outcomes. Some of the most impressive advances have been made with targeted therapies designed to treat cancers with specific causal mutations in critical genes known as driver genes.
These therapies have generally been developed to treat cancers where the driver gene has a “gain of function” mutation that causes cancer onset and progression. The goal of the therapies is to inhibit the newly acquired function of the gene itself or the pathways induced by the gene. Driver genes with a gain of function mutation, also known as oncogenes, are responsible for approximately 30% of all cancers. Most other cancers are driven by “loss of function” mutations in driver genes that normally suppress the onset of cancer. These driver genes are called tumor suppressor genes, and when mutated, they and their products cannot be targeted directly as their function is already turned off.
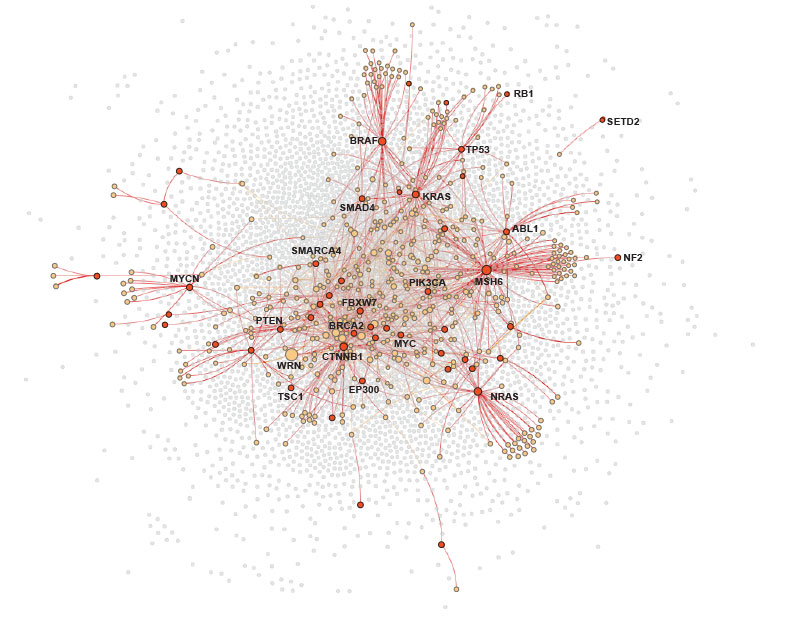
Accelerating Synthetic Lethality Insights
Synthetic lethality provides an avenue to treating cancers with loss of function mutations. A synthetic lethality relationship exists when a cell will die from a loss of function of two genes, but not from the loss of function of just one or the other. Since cancer cells already have mutated genes, a promising approach is to identify and then target pharmacologically a synthetically lethal partner of one of the mutated genes. Therapies utilizing this approach, such as PARP inhibitors, can successfully treat cancer while limiting toxicities in normal cells.
Leapfrogging Traditional SL Approaches
Leapfrog is redefining how genetically targeted therapies are discovered for cancers driven by loss-of-function mutations. Conventional synthetic-lethality approaches rely on genetic knockouts both to create the cancer-causing loss-of-function mutation and to evaluate potential drug targets. But drugs rarely replicate the effect of a knockout and often have clinically meaningful impacts that do not arise from a knockout. Our OncoSLX platform overcomes these limitations by using an advanced, deeply optimized pharmacogenetic strategy that evaluates the full biological effects of drugs for synergy with the same causal perturbations that drive human tumors.
Drugs do far more than eliminate a protein: they can block specific enzymatic activities, modulate unique functional domains, alter post-translational modifications present only in certain cellular states, or simultaneously inhibit multiple proteins. PARP inhibitors illustrate this principle—they synergize strongly with BRCA1 loss of function not because they mimic a PARP1 knockout, but because they trap PARP on DNA, a mechanism that is invisible to classic synthetic-lethality screens. By directly testing drug biology against true driver mutations, OncoSLX captures these pharmacological nuances and yields results that translate into patient benefit.
